Hello all, I come bearing plots to share my work on batching scipy.sparse.linalg.LinearOperator and its potential use in the iterative linear solvers.
Status
The linked PR is ready for review, and includes:
- batch support for the entire interface[1]
- tests across various kinds of operators and different kinds of batching
- a documentation overhaul
- housekeeping to reject
>2-Doperators in consuming algorithms for now - substantial self-review to aid reviewers
- green CI

I plan to let the PR sit for two to three weeks, before returning to hopefully make the final push to get it merged. Any reviews will be greatly appreciated!
Due diligence on un-batched performance
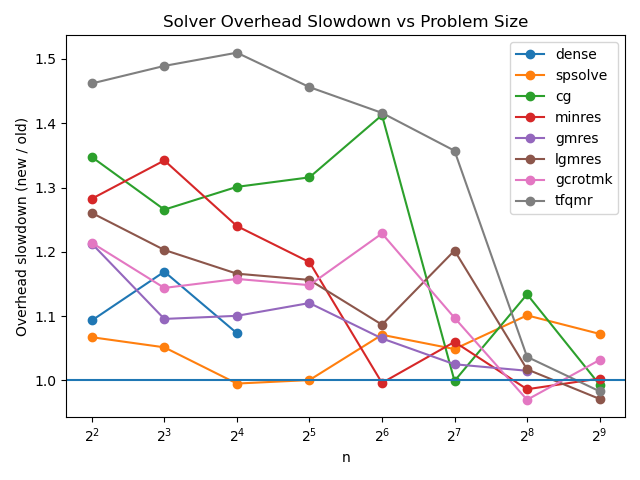
Firstly, I checked the overhead introduced for the un-batched case via the pre-existing benchmarks[2]:
The dense and spsolve lines don’t use the LinearOperator interface, so we can see that any additional overhead appears to disappear down to the level of noise by the time we get to n=2^8. The overhead for smaller problems seems sufficiently modest: the highest ratio comes from a <1ms delta, and the worst point there (tfqmr at n=2^7) is still just a delta of <20ms.
This seems very acceptable to me, although I don’t know exactly where the overhead is coming from, and why it seems to affect tfqmr more than the other solvers — anyone have a clue?
Batched cg performance
Now for the fun part: showing that this is useful. I have a rough implementation of batched conjugate gradient on the go at wip: N-D `LinearOperator` and `cg` by lucascolley · Pull Request #35 · lucascolley/scipy · GitHub which builds on top of the batched LinearOperator PR. This implementation supports batched LHS as well as the array API standard, but for now let me share some plots regarding performance with a single LHS and batched RHS.
The cg implementation is batched in the sense that it processes the entire input batch together, stepping all systems through each iteration simultaneously. Converged systems have to ‘go through the motions’ until every system has passed the convergence check. The advantage comes primarily from batched matvec calls.
n-D sparse.coo_array batched vs un-batched
For this benchmark, we use cg to solve (for \mathbf{x}) many instances of
where \mathbf{P} is Gilbert Strang’s favourite matrix of shape (n^2, n^2).
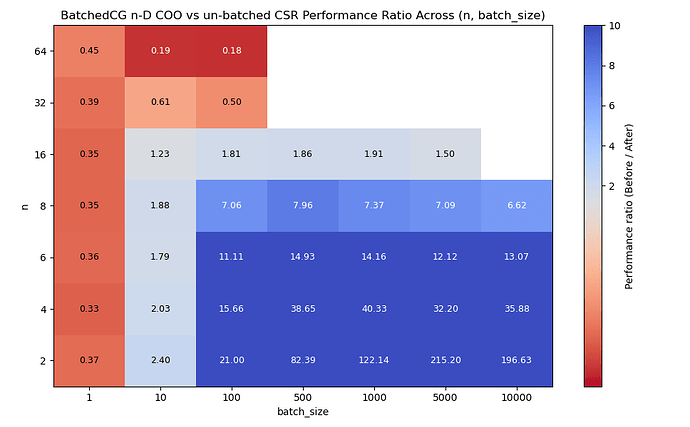
In the non-batched case, we represent \mathbf{P} as a scipy.sparse.csr_array[3], and solve against a list of b vectors in a Python for loop, where the list contains batch_size many vectors.
In the batched case, we represent the batch of \mathbf{P} as a 3-D scipy.sparse.coo_array of shape (batch_size, n*n, n*n), and b as a batch of vectors with shape (batch_size, n*n). We pass these to a single call of cg.
I ran the un-batched case on top of main to account for any new overhead. Here are the results[4]:
We see performance improvements of up to 200x for large batches of small systems.
Dense NumPy batched vs un-batched
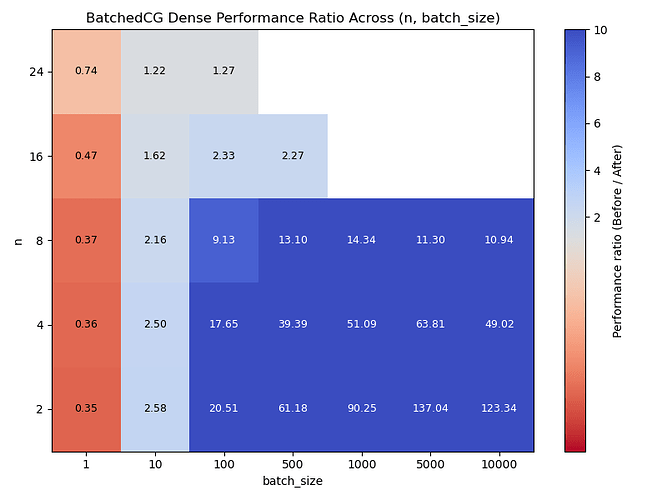
I also ran a benchmark with dense NumPy arrays. This time, for a matrix \mathbf{M} of shape (n^2, n^2) with normally distributed entries, we define a linear operator A by the function:
def matvec(x):
return np.squeeze(M.mT @ (M @ x[..., np.newaxis]), axis=-1) + 1e-3 * x
We use cg to solve (for \mathbf{x}) many instances of
We see similar results, perhaps performing notably better around n=8.
Overhead check
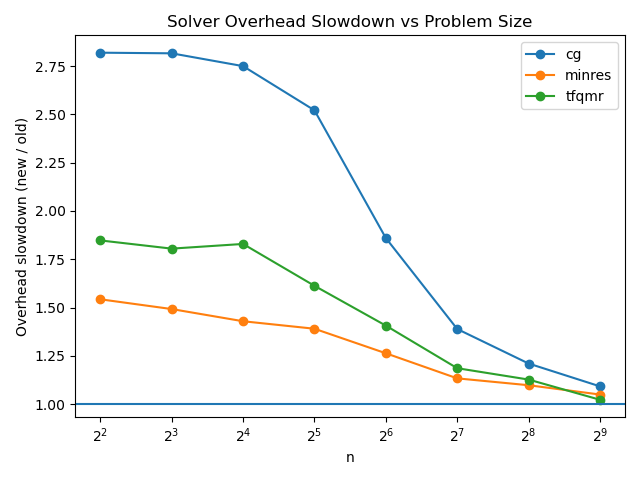
I also checked how much overhead the rough array API standard support implementation adds for the un-batched case:
The overhead is more prominent while we are down at the millisecond level, but it again falls to a seemingly acceptable level for sufficiently large matrices, so I don’t think this needs to worry us yet.
Further work
There is a lot of work left on the table, and I plan to spend the coming months investigating much of it, including:
- benchmarking with pathological cases in the batch
- benchmarking of batched LHS
- benchmarking with
pydata/sparse - benchmarking of array API standard support and on the GPU
- work on the other iterative solvers in
sparse.linalg - investigating more parallelism strategies (dividing batches in sub-batches?[5])
- investigating possible algorithmic changes, like ‘zeroing-out’ of rows corresponding to converged systems
Hopefully the results so far show that this is something worth pursuing!
It has been a fun but challenging exercise wrangling this section of code which had come to need some TLC as the years went by. I look forward to continuing the effort soon!
With happy holiday wishes ![]()
![]()
![]() ,
,
Lucas
P.S. for anyone wondering how on earth I’ve found the time to work on this, I am planning to write up my Master’s dissertation based on this work.
with one caveat related to the behaviour of an undocumented
__rmul__↩︎Results generated with
pixi run bench -t sparse_linalg_solve.Bench --compare base-bench-23836on bench · lucascolley/scipy@20a8769 · GitHub and bench · lucascolley/scipy@eb46853 · GitHub. Plot generated with the script at 14/12/2025 Batched LinearOperator overhead plot code · GitHub. ↩︎This was pre-existing in the benchmarks, I assume it is expected to be the fastest? In any case, I also benchmarked this against using COO arrays in the un-batched case and found very similar results. ↩︎
Results generated with
pixi run bench -t sparse_linalg_solve.BatchedCG --compare base-bench-23836on WIP: benchmarking · lucascolley/scipy@bc57a7c · GitHub and bench · lucascolley/scipy@eb46853 · GitHub. Plot generated with the script at … ↩︎Performance Portable Batched Sparse Linear Solvers | IEEE Journals & Magazine | IEEE Xplore takes that approach of dishing out mini-batches to ‘teams’, which seems to work harmoniously with the parallelism infrastructure available in Kokkos. ↩︎